Skull Stripping
Skull stripping is the process of removing the skull from brain MRI images to isolate the brain for further analysis.
Papers and Code
Reinforcing the Weakest Links: Modernizing SIENA with Targeted Deep Learning Integration
Mar 13, 2026Percentage Brain Volume Change (PBVC) derived from Magnetic Resonance Imaging (MRI) is a widely used biomarker of brain atrophy, with SIENA among the most established methods for its estimation. However, SIENA relies on classical image processing steps, particularly skull stripping and tissue segmentation, whose failures can propagate through the pipeline and bias atrophy estimates. In this work, we examine whether targeted deep learning substitutions can improve SIENA while preserving its established and interpretable framework. To this end, we integrate SynthStrip and SynthSeg into SIENA and evaluate three pipeline variants on the ADNI and PPMI longitudinal cohorts. Performance is assessed using three complementary criteria: correlation with longitudinal clinical and structural decline, scan-order consistency, and end-to-end runtime. Replacing the skull-stripping module yields the most consistent gains: in ADNI, it substantially strengthens associations between PBVC and multiple measures of disease progression relative to the standard SIENA pipeline, while across both datasets it markedly improves robustness under scan reversal. The fully integrated pipeline achieves the strongest scan-order consistency, reducing the error by up to 99.1%. In addition, GPU-enabled variants reduce execution time by up to 46% while maintaining CPU runtimes comparable to standard SIENA. Overall, these findings show that deep learning can meaningfully strengthen established longitudinal atrophy pipelines when used to reinforce their weakest image processing steps. More broadly, this study highlights the value of modularly modernizing clinically trusted neuroimaging tools without sacrificing their interpretability. Code is publicly available at https://github.com/Raciti/Enhanced-SIENA.git.
Simple Image Processing and Similarity Measures Can Link Data Samples across Databases through Brain MRI
Feb 10, 2026Head Magnetic Resonance Imaging (MRI) is routinely collected and shared for research under strict regulatory frameworks. These frameworks require removing potential identifiers before sharing. But, even after skull stripping, the brain parenchyma contains unique signatures that can match other MRIs from the same participants across databases, posing a privacy risk if additional data features are available. Current regulatory frameworks often mandate evaluating such risks based on the assessment of a certain level of reasonableness. Prior studies have already suggested that a brain MRI could enable participant linkage, but they have relied on training-based or computationally intensive methods. Here, we demonstrate that linking an individual's skull-stripped T1-weighted MRI, which may lead to re-identification if other identifiers are available, is possible using standard preprocessing followed by image similarity computation. Nearly perfect linkage accuracy was achieved in matching data samples across various time intervals, scanner types, spatial resolutions, and acquisition protocols, despite potential cognitive decline, simulating MRI matching across databases. These results aim to contribute meaningfully to the development of thoughtful, forward-looking policies in medical data sharing.
Efficient Brain Extraction of MRI Scans with Mild to Moderate Neuropathology
Feb 09, 2026Skull stripping magnetic resonance images (MRI) of the human brain is an important process in many image processing techniques, such as automatic segmentation of brain structures. Numerous methods have been developed to perform this task, however, they often fail in the presence of neuropathology and can be inconsistent in defining the boundary of the brain mask. Here, we propose a novel approach to skull strip T1-weighted images in a robust and efficient manner, aiming to consistently segment the outer surface of the brain, including the sulcal cerebrospinal fluid (CSF), while excluding the full extent of the subarachnoid space and meninges. We train a modified version of the U-net on silver-standard ground truth data using a novel loss function based on the signed-distance transform (SDT). We validate our model both qualitatively and quantitatively using held-out data from the training dataset, as well as an independent external dataset. The brain masks used for evaluation partially or fully include the subarachnoid space, which may introduce bias into the comparison; nonetheless, our model demonstrates strong performance on the held-out test data, achieving a consistent mean Dice similarity coefficient (DSC) of 0.964$\pm$0.006 and an average symmetric surface distance (ASSD) of 1.4mm$\pm$0.2mm. Performance on the external dataset is comparable, with a DSC of 0.958$\pm$0.006 and an ASSD of 1.7$\pm$0.2mm. Our method achieves performance comparable to or better than existing state-of-the-art methods for brain extraction, particularly in its highly consistent preservation of the brain's outer surface. The method is publicly available on GitHub.
MindGrab for BrainChop: Fast and Accurate Skull Stripping for Command Line and Browser
Jun 13, 2025We developed MindGrab, a parameter- and memory-efficient deep fully-convolutional model for volumetric skull-stripping in head images of any modality. Its architecture, informed by a spectral interpretation of dilated convolutions, was trained exclusively on modality-agnostic synthetic data. MindGrab was evaluated on a retrospective dataset of 606 multimodal adult-brain scans (T1, T2, DWI, MRA, PDw MRI, EPI, CT, PET) sourced from the SynthStrip dataset. Performance was benchmarked against SynthStrip, ROBEX, and BET using Dice scores, with Wilcoxon signed-rank significance tests. MindGrab achieved a mean Dice score of 95.9 with standard deviation (SD) 1.6 across modalities, significantly outperforming classical methods (ROBEX: 89.1 SD 7.7, P < 0.05; BET: 85.2 SD 14.4, P < 0.05). Compared to SynthStrip (96.5 SD 1.1, P=0.0352), MindGrab delivered equivalent or superior performance in nearly half of the tested scenarios, with minor differences (<3% Dice) in the others. MindGrab utilized 95% fewer parameters (146,237 vs. 2,566,561) than SynthStrip. This efficiency yielded at least 2x faster inference, 50% lower memory usage on GPUs, and enabled exceptional performance (e.g., 10-30x speedup, and up to 30x memory reduction) and accessibility on a wider range of hardware, including systems without high-end GPUs. MindGrab delivers state-of-the-art accuracy with dramatically lower resource demands, supported in brainchop-cli (https://pypi.org/project/brainchop/) and at brainchop.org.
Skull stripping with purely synthetic data
May 12, 2025While many skull stripping algorithms have been developed for multi-modal and multi-species cases, there is still a lack of a fundamentally generalizable approach. We present PUMBA(PUrely synthetic Multimodal/species invariant Brain extrAction), a strategy to train a model for brain extraction with no real brain images or labels. Our results show that even without any real images or anatomical priors, the model achieves comparable accuracy in multi-modal, multi-species and pathological cases. This work presents a new direction of research for any generalizable medical image segmentation task.
How We Won the ISLES'24 Challenge by Preprocessing
May 23, 2025

Stroke is among the top three causes of death worldwide, and accurate identification of stroke lesion boundaries is critical for diagnosis and treatment. Supervised deep learning methods have emerged as the leading solution for stroke lesion segmentation but require large, diverse, and annotated datasets. The ISLES'24 challenge addresses this need by providing longitudinal stroke imaging data, including CT scans taken on arrival to the hospital and follow-up MRI taken 2-9 days from initial arrival, with annotations derived from follow-up MRI. Importantly, models submitted to the ISLES'24 challenge are evaluated using only CT inputs, requiring prediction of lesion progression that may not be visible in CT scans for segmentation. Our winning solution shows that a carefully designed preprocessing pipeline including deep-learning-based skull stripping and custom intensity windowing is beneficial for accurate segmentation. Combined with a standard large residual nnU-Net architecture for segmentation, this approach achieves a mean test Dice of 28.5 with a standard deviation of 21.27.
DISARM++: Beyond scanner-free harmonization
May 06, 2025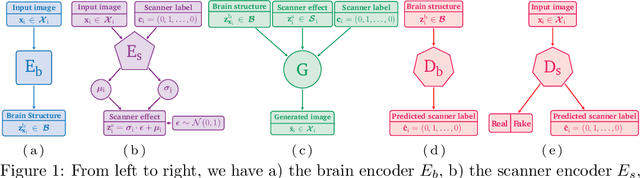
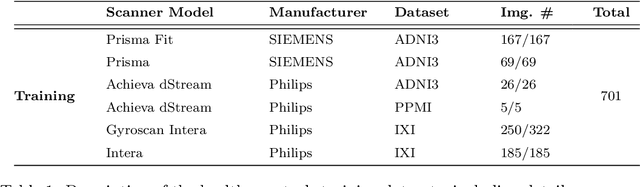
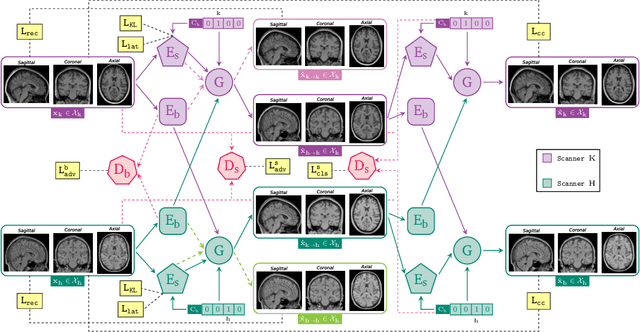
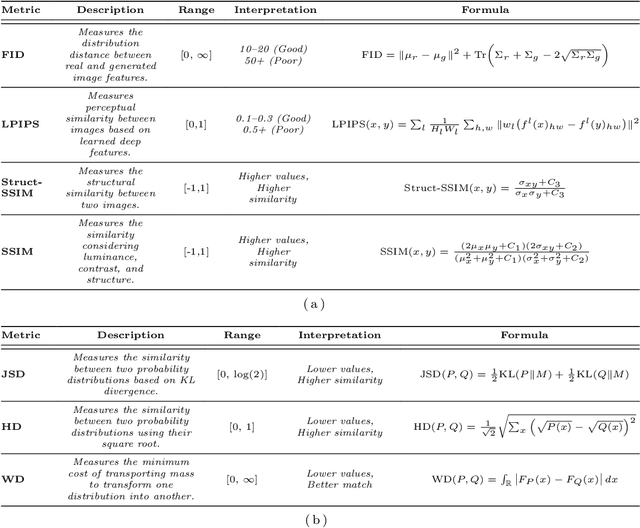
Harmonization of T1-weighted MR images across different scanners is crucial for ensuring consistency in neuroimaging studies. This study introduces a novel approach to direct image harmonization, moving beyond feature standardization to ensure that extracted features remain inherently reliable for downstream analysis. Our method enables image transfer in two ways: (1) mapping images to a scanner-free space for uniform appearance across all scanners, and (2) transforming images into the domain of a specific scanner used in model training, embedding its unique characteristics. Our approach presents strong generalization capability, even for unseen scanners not included in the training phase. We validated our method using MR images from diverse cohorts, including healthy controls, traveling subjects, and individuals with Alzheimer's disease (AD). The model's effectiveness is tested in multiple applications, such as brain age prediction (R2 = 0.60 \pm 0.05), biomarker extraction, AD classification (Test Accuracy = 0.86 \pm 0.03), and diagnosis prediction (AUC = 0.95). In all cases, our harmonization technique outperforms state-of-the-art methods, showing improvements in both reliability and predictive accuracy. Moreover, our approach eliminates the need for extensive preprocessing steps, such as skull-stripping, which can introduce errors by misclassifying brain and non-brain structures. This makes our method particularly suitable for applications that require full-head analysis, including research on head trauma and cranial deformities. Additionally, our harmonization model does not require retraining for new datasets, allowing smooth integration into various neuroimaging workflows. By ensuring scanner-invariant image quality, our approach provides a robust and efficient solution for improving neuroimaging studies across diverse settings. The code is available at this link.
PhaseGen: A Diffusion-Based Approach for Complex-Valued MRI Data Generation
Apr 10, 2025Magnetic resonance imaging (MRI) raw data, or k-Space data, is complex-valued, containing both magnitude and phase information. However, clinical and existing Artificial Intelligence (AI)-based methods focus only on magnitude images, discarding the phase data despite its potential for downstream tasks, such as tumor segmentation and classification. In this work, we introduce $\textit{PhaseGen}$, a novel complex-valued diffusion model for generating synthetic MRI raw data conditioned on magnitude images, commonly used in clinical practice. This enables the creation of artificial complex-valued raw data, allowing pretraining for models that require k-Space information. We evaluate PhaseGen on two tasks: skull-stripping directly in k-Space and MRI reconstruction using the publicly available FastMRI dataset. Our results show that training with synthetic phase data significantly improves generalization for skull-stripping on real-world data, with an increased segmentation accuracy from $41.1\%$ to $80.1\%$, and enhances MRI reconstruction when combined with limited real-world data. This work presents a step forward in utilizing generative AI to bridge the gap between magnitude-based datasets and the complex-valued nature of MRI raw data. This approach allows researchers to leverage the vast amount of avaliable image domain data in combination with the information-rich k-Space data for more accurate and efficient diagnostic tasks. We make our code publicly $\href{https://github.com/TIO-IKIM/PhaseGen}{\text{available here}}$.
Pfungst and Clever Hans: Identifying the unintended cues in a widely used Alzheimer's disease MRI dataset using explainable deep learning
Jan 27, 2025



Backgrounds. Deep neural networks have demonstrated high accuracy in classifying Alzheimer's disease (AD). This study aims to enlighten the underlying black-box nature and reveal individual contributions of T1-weighted (T1w) gray-white matter texture, volumetric information and preprocessing on classification performance. Methods. We utilized T1w MRI data from the Alzheimer's Disease Neuroimaging Initiative to distinguish matched AD patients (990 MRIs) from healthy controls (990 MRIs). Preprocessing included skull stripping and binarization at varying thresholds to systematically eliminate texture information. A deep neural network was trained on these configurations, and the model performance was compared using McNemar tests with discrete Bonferroni-Holm correction. Layer-wise Relevance Propagation (LRP) and structural similarity metrics between heatmaps were applied to analyze learned features. Results. Classification performance metrics (accuracy, sensitivity, and specificity) were comparable across all configurations, indicating a negligible influence of T1w gray- and white signal texture. Models trained on binarized images demonstrated similar feature performance and relevance distributions, with volumetric features such as atrophy and skull-stripping features emerging as primary contributors. Conclusions. We revealed a previously undiscovered Clever Hans effect in a widely used AD MRI dataset. Deep neural networks classification predominantly rely on volumetric features, while eliminating gray-white matter T1w texture did not decrease the performance. This study clearly demonstrates an overestimation of the importance of gray-white matter contrasts, at least for widely used structural T1w images, and highlights potential misinterpretation of performance metrics.
Unsupervised Skull Segmentation via Contrastive MR-to-CT Modality Translation
Oct 17, 2024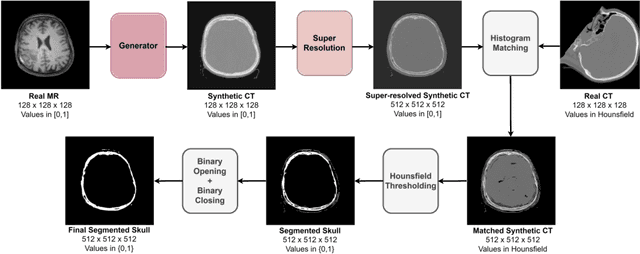
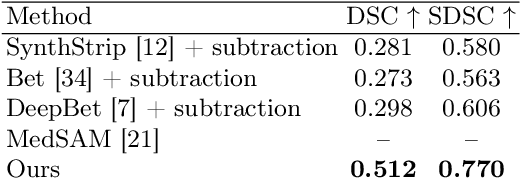

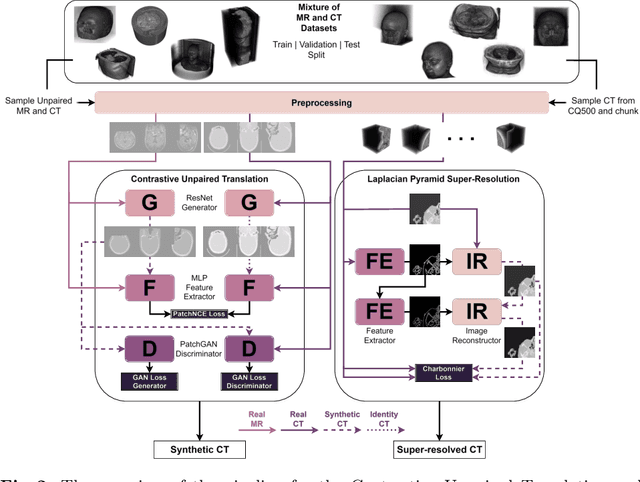
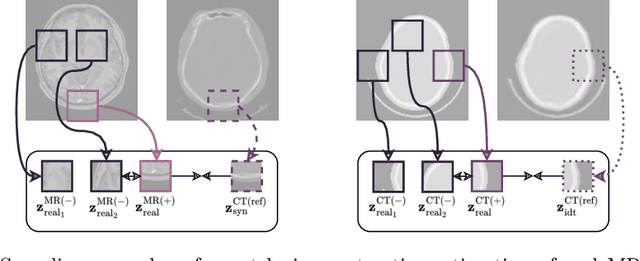
The skull segmentation from CT scans can be seen as an already solved problem. However, in MR this task has a significantly greater complexity due to the presence of soft tissues rather than bones. Capturing the bone structures from MR images of the head, where the main visualization objective is the brain, is very demanding. The attempts that make use of skull stripping seem to not be well suited for this task and fail to work in many cases. On the other hand, supervised approaches require costly and time-consuming skull annotations. To overcome the difficulties we propose a fully unsupervised approach, where we do not perform the segmentation directly on MR images, but we rather perform a synthetic CT data generation via MR-to-CT translation and perform the segmentation there. We address many issues associated with unsupervised skull segmentation including the unpaired nature of MR and CT datasets (contrastive learning), low resolution and poor quality (super-resolution), and generalization capabilities. The research has a significant value for downstream tasks requiring skull segmentation from MR volumes such as craniectomy or surgery planning and can be seen as an important step towards the utilization of synthetic data in medical imaging.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge